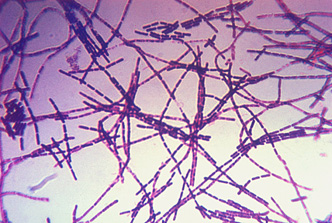
A photomicrograph of Bacillus anthracis bacteria using Gram stain technique.
Anthrax is diagnosed by isolating B. anthracis from the blood, skin lesions, or respiratory secretions or by measuring specific antibodies in the blood of persons with suspected cases.
By following a DNA trail through the secretive world of heroin use, researchers may have shown the way for public health officials to solve infectious disease problems affecting the wider population.
Led by Paul Keim, Regents’ Professor and Cowden Endowed Chair of Microbiology at Northern Arizona University, a collaborative team of international scientists shone new light on an old question about injectional anthrax by demonstrating the latest advances in whole genome sequencing. Researchers from Germany, England and the United States reconstructed a decades-old outbreak of injectional anthrax by DNA analysis of the anthrax conveyed to victims from injecting heroin.
The results were pre-published online at EBioMedicine in advance of their appearance in the final version of the journal. In the article, the research team demonstrated the potential of whole genome sequencing, a data-rich process used to determine an organism’s entire DNA sequence.
“It’s an application of a technology that’s been incrementally improving over the last decade,” said Keim, who used a similar approach to the anthrax letter attack in a case that garnered widespread attention amidst fears of bioterrorism. But today’s sequencing work is far more expansive, rapid and less expensive—so much so that the insights from this project have added new considerations to some of the foundational principles of epidemiology.
“This work is an example of the paradigm for future investigations,” Keim said.
The problem being investigated—anthrax infections resulting from the injection of heroin—was the topic of analysis three years ago by the Center for Microbial Genetics and Genomics, an NAU lab directed by Keim. At that time, evidence pointed to the idea that heroin had become contaminated by anthrax along the way from Afghanistan to Europe, where some heroin users were becoming infected and even dying.
While the new investigation leaves open the question of where the contamination took place, it identified two distinct disease clusters occurring over a period of more than 12 years. The conclusion defied the epidemiological bedrock of space-time correlation that had pointed to only one disease event, Keim said.
The implications are powerful for public health officials who face obstacles of money and time to accurately identify and track a disease event, such as a food-borne pathogen outbreak.
“That’s the whole purpose of doing molecular epidemiology,” Keim said. “And this is the biggest, baddest way to do it.”
NAU played a central role in the international effort because of its inventory of more than 3,000 anthrax strains and because of its investment in high-capacity computation, known on campus as the Monsoon computing cluster. Keim said some of the data files transferred to his lab were so large they took weeks to process, even with Monsoon.
But Keim pointed to the exchange of information as an example of how whole genome sequencing effectively crosses international boundaries, which restrict the distribution of live anthrax.
“This type of technology is very amenable to standardization,” Keim said, noting that this project involved two labs in Germany, two in the United Kingdom, the U.S. Centers for Disease Control and NAU. “You can easily put genome sequences into a standard language for the exchange of data. It’s like putting a pie back together if you have all the slices.”
Keim said one of the goals of the research funding from the U.S. Department of Homeland Security was to encourage such global collaboration.
“They wanted international scientists to use this technology to solve a problem and establish ties,” Keim said. “If there is ever such an event, we have established the international cooperation for an investigation. This is setting the stage for future collaborations.”
Source: http://news.nau.edu/naus-lead-in-anthrax-research-sets-tone-for-international-public-health-investigations/